Data Visualization with ggplot2
Last updated on 2026-03-31 | Edit this page
Estimated time: 90 minutes
Overview
Questions
- What is ggplot2?
- What is mapping, and what is aesthetics?
- What is the process of creating a publication-quality plots with ggplot in R?
Objectives
- Describe the role of data, aesthetics, and geoms in ggplot functions.
- Choose the correct aesthetics and alter the geom parameters for a scatter plot, histogram, or box plot.
- Layer multiple geometries in a single plot.
- Customize plot scales, titles, themes, and fonts.
- Apply a facet to a plot.
- Apply additional ggplot2-compatible plotting libraries.
- Save a ggplot to a file.
- List several resources for getting help with ggplot.
- List several resources for creating informative scientific plots.
Reminder
At this point you should be coding along in the “genomics_r_plotting.R” script we created in the last episode. Writing your commands in the script (and commenting it) will make it easier to record what you did and why.
Introduction to ggplot2

ggplot2 is a plotting package, part of
the tidyverse, that makes it simple to create complex plots from data in
a data frame. It provides a more programmatic interface for specifying
what variables to plot, how they are displayed, and general visual
properties. Therefore, we only need minimal changes if the underlying
data change or if we decide to change from a bar plot to a scatter plot.
This helps in creating publication-quality plots with minimal amounts of
adjustments and tweaking.
The gg in “ggplot” stands for “Grammar of Graphics,” which is an elegant yet powerful way to describe the making of scientific plots. In short, the grammar of graphics breaks down every plot into a few components, namely, a dataset, a set of geoms (visual marks that represent the data points), and a coordinate system. You can imagine this is a grammar that gives unique names to each component appearing in a plot and conveys specific information about data. With ggplot, graphics are built step by step by adding new elements.
The idea of mapping is crucial in
ggplot. One familiar example is to map the
value of one variable in a dataset to \(x\) and the other to \(y\). However, we often encounter datasets
that include multiple (more than two) variables. In this case,
ggplot allows you to map those other variables
to visual marks such as color and
shape (aesthetics or
aes). One thing you may want to remember is the difference
between discrete and continuous
variables. Some aesthetics, such as the shape of dots, do not accept
continuous variables. If forced to do so, R will give an error. This is
easy to understand; we cannot create a continuum of shapes for a
variable, unlike, say, color.
Tip: when having doubts about whether a variable is
continuous
or discrete, a quick way to check is to use the summary()
function. Continuous variables have descriptive statistics but not the
discrete variables.
Installing and loading ggplot
In your
“genomics_r"genomics_r_basics.R”.R”
script type the following code to load the ggplot2
package.
R
library(ggplot2)
Next, run this line of code in your script. You can run a line of code by hitting the Run button that is just above the first line of your script in the header of the Source pane or you can use the appropriate shortcut:
- Windows execution shortcut: Ctrl+Enter
- Mac execution shortcut: Cmd(⌘)+Enter
What do you see as an output in the Console? Do you see an error that reads Error in library(ggplot2) : there is no package called ‘ggplot2’? If so, this means you don’t have this external third-party library installed. We’ll talk later about external libraries, but for now we’ll just focus on having it installed to use it.
If you got the error, type the following code in the Console to
install the ggplot2 package.
R
install.packages("ggplot2")
Loading the dataset
R
variants <- read.csv("https://www.tinyurl.com/r-sddrc-data")
One of the first things you should notice is that in the Environment
window, you have the variants object, listed as 801 obs.
(observations/rows) of 29 variables (columns). Double-clicking on the
name of the object will open a view of the data in a new tab.
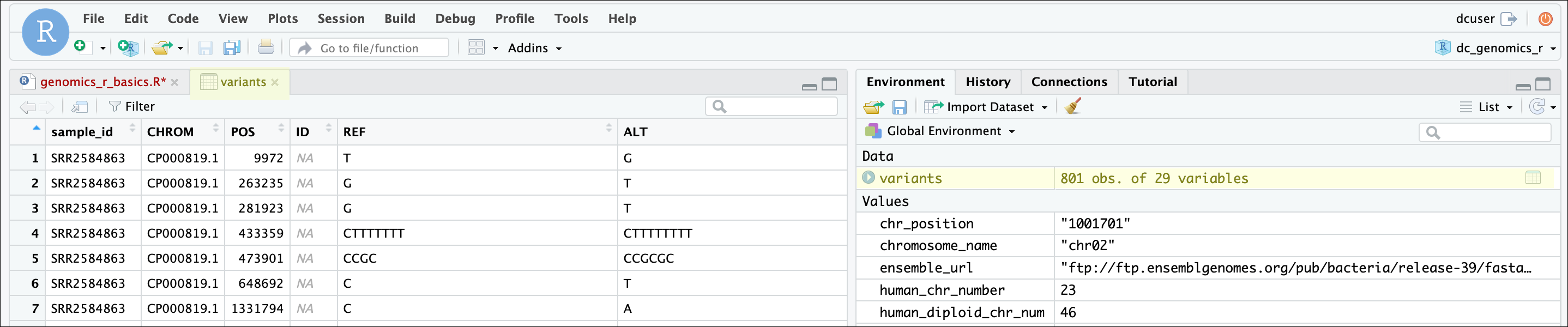
Explore the structure (types of columns and number of rows)
of the dataset using the str() function.
R
str(variants) # Show the structure of the data
OUTPUT
'data.frame': 801 obs. of 29 variables:
$ sample_id : chr "SRR2584863" "SRR2584863" "SRR2584863" "SRR2584863" ...
$ CHROM : chr "CP000819.1" "CP000819.1" "CP000819.1" "CP000819.1" ...
$ POS : int 9972 263235 281923 433359 473901 648692 1331794 1733343 2103887 2333538 ...
$ ID : logi NA NA NA NA NA NA ...
$ REF : chr "T" "G" "G" "CTTTTTTT" ...
$ ALT : chr "G" "T" "T" "CTTTTTTTT" ...
$ QUAL : num 91 85 217 64 228 210 178 225 56 167 ...
$ FILTER : logi NA NA NA NA NA NA ...
$ INDEL : logi FALSE FALSE FALSE TRUE TRUE FALSE ...
$ IDV : int NA NA NA 12 9 NA NA NA 2 7 ...
$ IMF : num NA NA NA 1 0.9 ...
$ DP : int 4 6 10 12 10 10 8 11 3 7 ...
$ VDB : num 0.0257 0.0961 0.7741 0.4777 0.6595 ...
$ RPB : num NA 1 NA NA NA NA NA NA NA NA ...
$ MQB : num NA 1 NA NA NA NA NA NA NA NA ...
$ BQB : num NA 1 NA NA NA NA NA NA NA NA ...
$ MQSB : num NA NA 0.975 1 0.916 ...
$ SGB : num -0.556 -0.591 -0.662 -0.676 -0.662 ...
$ MQ0F : num 0 0.167 0 0 0 ...
$ ICB : logi NA NA NA NA NA NA ...
$ HOB : logi NA NA NA NA NA NA ...
$ AC : int 1 1 1 1 1 1 1 1 1 1 ...
$ AN : int 1 1 1 1 1 1 1 1 1 1 ...
$ DP4 : chr "0,0,0,4" "0,1,0,5" "0,0,4,5" "0,1,3,8" ...
$ MQ : int 60 33 60 60 60 60 60 60 60 60 ...
$ Indiv : chr "/home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam" "/home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam" "/home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam" "/home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam" ...
$ gt_PL : chr "121,0" "112,0" "247,0" "91,0" ...
$ gt_GT : int 1 1 1 1 1 1 1 1 1 1 ...
$ gt_GT_alleles: chr "G" "T" "T" "CTTTTTTTT" ...To run multiple lines of code, you can highlight all the line you wish to run and then hit Run or use the shortcut key combo listed above.
Alternatively, we can display the first a few rows (vertically) of
the table using head():
R
head(variants)
| sample_id | CHROM | POS | ID | REF | ALT | QUAL | FILTER | INDEL | IDV | IMF | DP | VDB | RPB | MQB | BQB | MQSB | SGB | MQ0F | ICB | HOB | AC | AN | DP4 | MQ | Indiv | gt_PL | gt_GT | gt_GT_alleles |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SRR2584863 | CP000819.1 | 9972 | NA | T | G | 91 | NA | FALSE | NA | NA | 4 | 0.0257451 | NA | NA | NA | NA | -0.556411 | 0.000000 | NA | NA | 1 | 1 | 0,0,0,4 | 60 | /home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam | 121,0 | 1 | G |
| SRR2584863 | CP000819.1 | 263235 | NA | G | T | 85 | NA | FALSE | NA | NA | 6 | 0.0961330 | 1 | 1 | 1 | NA | -0.590765 | 0.166667 | NA | NA | 1 | 1 | 0,1,0,5 | 33 | /home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam | 112,0 | 1 | T |
| SRR2584863 | CP000819.1 | 281923 | NA | G | T | 217 | NA | FALSE | NA | NA | 10 | 0.7740830 | NA | NA | NA | 0.974597 | -0.662043 | 0.000000 | NA | NA | 1 | 1 | 0,0,4,5 | 60 | /home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam | 247,0 | 1 | T |
| SRR2584863 | CP000819.1 | 433359 | NA | CTTTTTTT | CTTTTTTTT | 64 | NA | TRUE | 12 | 1.0 | 12 | 0.4777040 | NA | NA | NA | 1.000000 | -0.676189 | 0.000000 | NA | NA | 1 | 1 | 0,1,3,8 | 60 | /home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam | 91,0 | 1 | CTTTTTTTT |
| SRR2584863 | CP000819.1 | 473901 | NA | CCGC | CCGCGC | 228 | NA | TRUE | 9 | 0.9 | 10 | 0.6595050 | NA | NA | NA | 0.916482 | -0.662043 | 0.000000 | NA | NA | 1 | 1 | 1,0,2,7 | 60 | /home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam | 255,0 | 1 | CCGCGC |
| SRR2584863 | CP000819.1 | 648692 | NA | C | T | 210 | NA | FALSE | NA | NA | 10 | 0.2680140 | NA | NA | NA | 0.916482 | -0.670168 | 0.000000 | NA | NA | 1 | 1 | 0,0,7,3 | 60 | /home/dcuser/dc_workshop/results/bam/SRR2584863.aligned.sorted.bam | 240,0 | 1 | T |
We can also see view our data in a nicely formatted window using the
View() function, which opens a new tab in our Source pane.
This will have the same result as clicking the name of our dataset in
the Environment pane.
R
View(variants)
ggplot2 functions like data in the
long format, i.e., a column for every dimension
(variable), and a row for every observation. Well-structured data will
save you time when making figures with
ggplot2
ggplot2 graphics are built step-by-step
by adding new elements. Adding layers in this fashion allows for
extensive flexibility and customization of plots, and more equally
important the readability of the code.
To build a ggplot, we will use the following basic template that can be used for different types of plots:
- use the
ggplot()function and bind the plot to a specific data frame using thedataargument
R
ggplot(data = variants)
- define a mapping (using the aesthetic (
aes) function), by selecting the variables to be plotted and specifying how to present them in the graph, e.g. as x and y positions or characteristics such as size, shape, color, etc.
R
ggplot(data = variants, aes(x = DP, y = QUAL))
- add ‘geoms’ – graphical representations of the data in the plot
(points, lines, bars).
ggplot2offers many different geoms; we will use some common ones today, including:-
geom_point()for scatter plots, dot plots, etc. -
geom_boxplot()for, well, boxplots! -
geom_line()for trend lines, time series, etc.
-
To add a geom to the plot use the + operator. Because we
have two continuous variables, let’s use geom_point()
(i.e., a scatter plot) first:
R
ggplot(data = variants, aes(x = DP, y = QUAL)) +
geom_point()
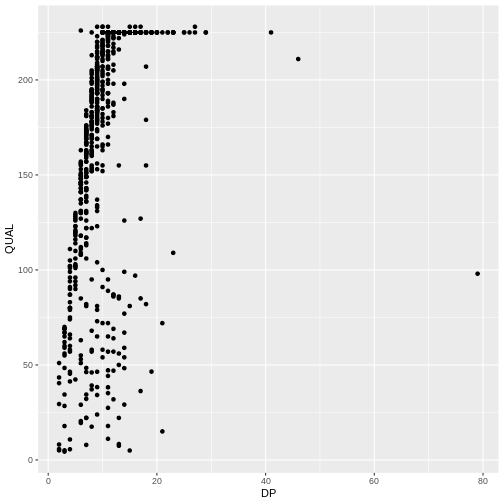
The + in the ggplot2
package is particularly useful because it allows you to modify existing
ggplot objects. This means you can easily set up plot
templates and conveniently explore different types of plots, so the
above plot can also be generated with code like this:
R
# Assign plot to a variable
coverage_plot <- ggplot(data = variants, aes(x = DP, y = QUAL))
# Draw the plot
coverage_plot +
geom_point()
Notes
- Anything you put in the
ggplot()function can be seen by any geom layers that you add (i.e., these are universal plot settings). This includes the x- and y-axis mapping you set up inaes(). - You can also specify mappings for a given geom independently of the
mappings defined globally in the
ggplot()function. - The
+sign used to add new layers must be placed at the end of the line containing the previous layer. If, instead, the+sign is added at the beginning of the line containing the new layer,ggplot2will not add the new layer and will return an error message.
R
# This is the correct syntax for adding layers
coverage_plot +
geom_point()
# This will not add the new layer and will return an error message
coverage_plot
+ geom_point()
Building your plots iteratively
Building plots with ggplot2 is
typically an iterative process. We start by defining the dataset we’ll
use, lay out the axes, and choose a geom:
R
ggplot(data = variants, aes(x = DP, y = QUAL)) +
geom_point()
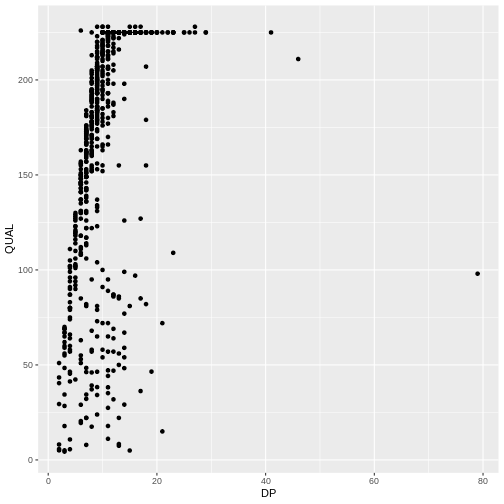
Then, we start modifying this plot to extract more information from
it. For instance, we can add transparency (alpha) to avoid
over-plotting:
R
ggplot(data = variants, aes(x = DP, y = QUAL)) +
geom_point(alpha = 0.5)
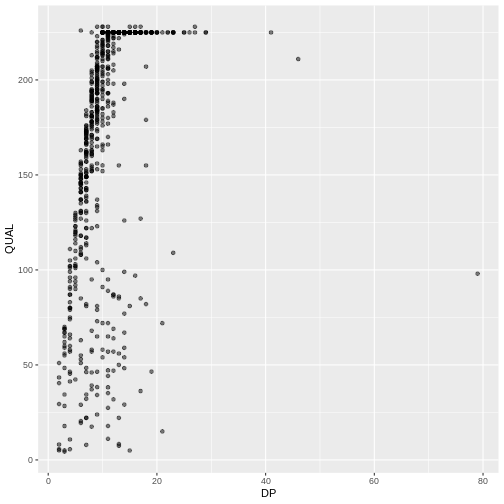
We can also add colors for all the points:
R
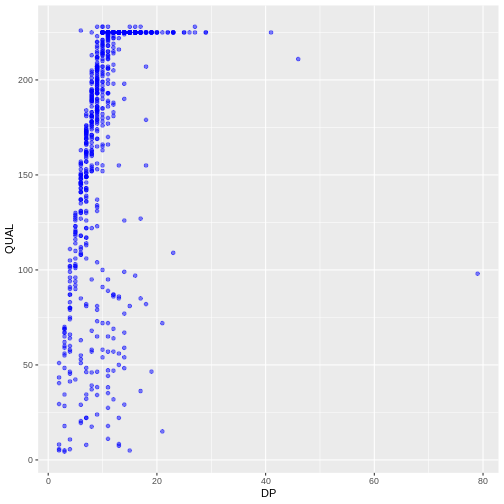
ggplot(data = variants, aes(x = DP, y = QUAL)) +
geom_point(alpha = 0.5, color = "blue")
Or to color each species in the plot differently, you could use a
vector as an input to the argument color.
ggplot2 will provide a different color
corresponding to different values in the vector. Here is an example
where we color with sample_id:
R
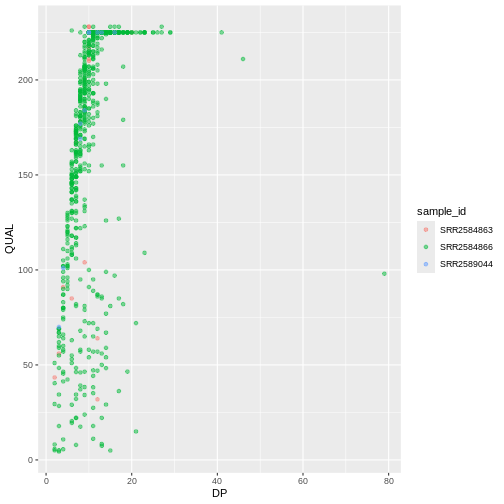
ggplot(data = variants, aes(x = DP, y = QUAL, color = sample_id)) +
geom_point(alpha = 0.5)
To make our plot more readable, we can add axis labels:
R
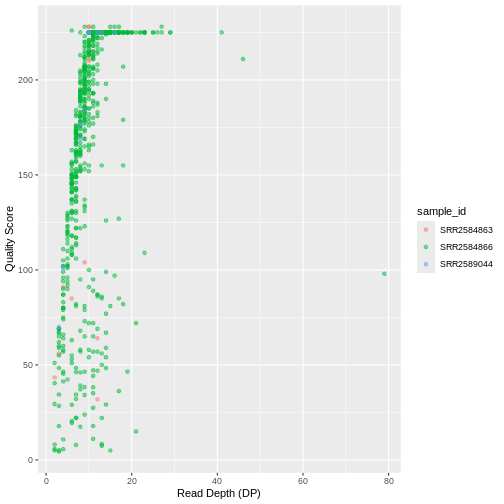
ggplot(data = variants, aes(x = DP, y = QUAL, color = sample_id)) +
geom_point(alpha = 0.5) +
labs(x = "Read Depth (DP)",
y = "Quality Score")
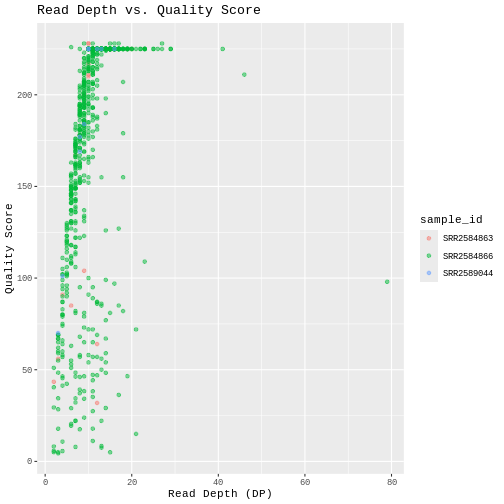
To add a main title to the plot, we use the title
argument for the labs() function:
R
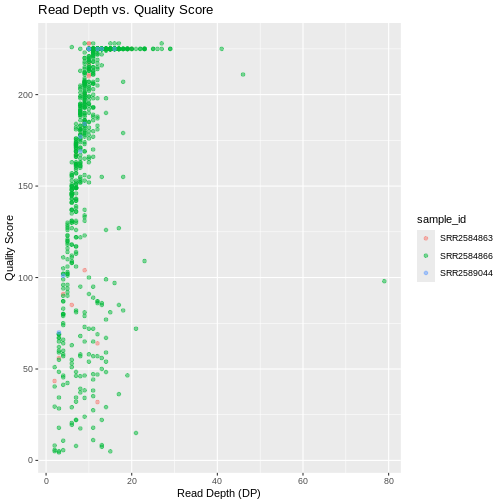
ggplot(data = variants, aes(x = DP, y = QUAL, color = sample_id)) +
geom_point(alpha = 0.5) +
labs(x = "Read Depth (DP)",
y = "Quality Score",
title = "Read Depth vs. Quality Score")
Now the figure is complete and ready to be exported and saved to a
file. This can be achieved easily using ggsave(),
which can write, by default, the most recent generated figure into
different formats (e.g., jpeg, png,
pdf) according to the file extension. So, for example, to
create a pdf version of the above figure with a dimension of \(6\times4\) inches:
R
ggsave("depth_quality.pdf", width = 6, height = 4)
If we check the current working directory, there should be a
newly created file called depth.pdf with the above
plot.
Challenge
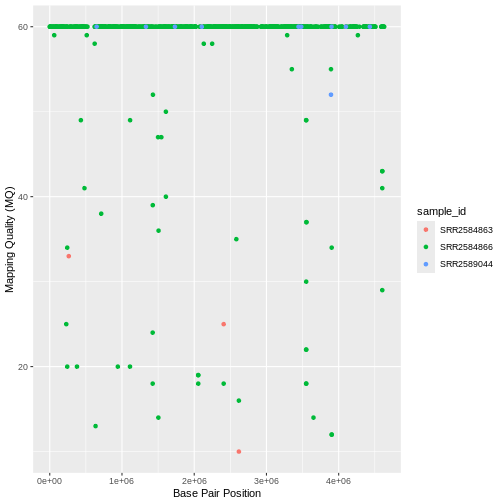
Use what you just learned to create a scatter plot of mapping quality
(MQ) over position (POS) with the samples
showing in different colors. Make sure to give your plot relevant axis
labels.
R
ggplot(data = variants, aes(x = POS, y = MQ, color = sample_id)) +
geom_point() +
labs(x = "Base Pair Position",
y = "Mapping Quality (MQ)")
To further customize the plot, we can change the default font format:
R
ggplot(data = variants, aes(x = DP, y = QUAL, color = sample_id)) +
geom_point(alpha = 0.5) +
labs(x = "Read Depth (DP)",
y = "Quality Score",
title = "Read Depth vs. Quality Score") +
theme(text = element_text(family = "mono"))
Faceting
ggplot2 has a special technique called
faceting that allows the user to split one plot into multiple
plots (panels) based on a factor (variable) included in the dataset. We
will use it to split our mapping quality plot into three panels, one for
each sample.
R
ggplot(data = variants, aes(x = DP, y = QUAL)) +
geom_point(alpha = 0.5) +
labs(x = "Read Depth (DP)",
y = "Quality Score",
title = "Read Depth vs. Quality Score") +
facet_grid(~ sample_id)
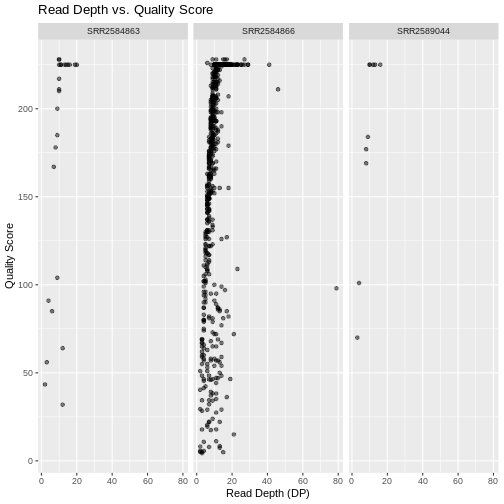
This looks okay, but it would be easier to read if the plot facets
were stacked vertically rather than horizontally. The
facet_grid geometry allows you to explicitly specify how
you want your plots to be arranged via formula notation
(rows ~ columns; the dot (.) indicates every
other variable in the data i.e., no faceting on that side of the
formula).
R
ggplot(data = variants, aes(x = DP, y = QUAL)) +
geom_point(alpha = 0.5) +
labs(x = "Read Depth (DP)",
y = "Quality Score",
title = "Read Depth vs. Quality Score") +
facet_grid(sample_id ~ .)
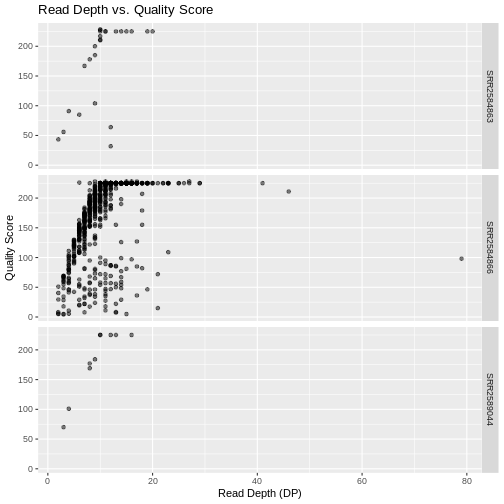
Usually plots with white background look more readable when printed.
We can set the background to white using the function theme_bw().
Additionally, you can remove the grid:
R
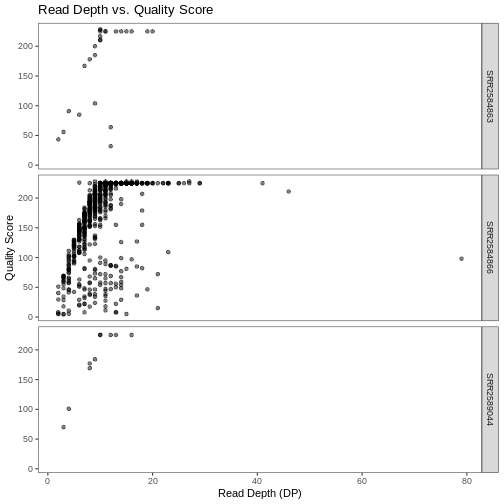
ggplot(data = variants, aes(x = DP, y = QUAL)) +
geom_point(alpha = 0.5) +
labs(x = "Read Depth (DP)",
y = "Quality Score",
title = "Read Depth vs. Quality Score") +
facet_grid(sample_id ~ .) +
theme_bw() +
theme(panel.grid = element_blank())
Challenge
Use what you just learned to create a scatter plot of PHRED scaled
quality (QUAL) over position (POS) with the
samples showing in different facets. Make sure to give your plot
relevant axis labels.
R
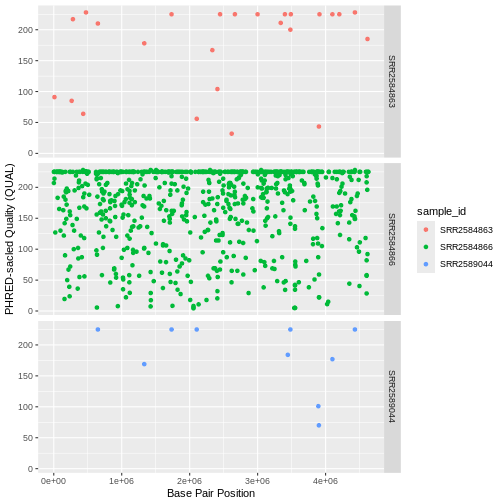
ggplot(data = variants, aes(x = POS, y = QUAL, color = sample_id)) +
geom_point() +
labs(x = "Base Pair Position",
y = "PHRED-sacled Quality (QUAL)") +
facet_grid(sample_id ~ .)
Barplots
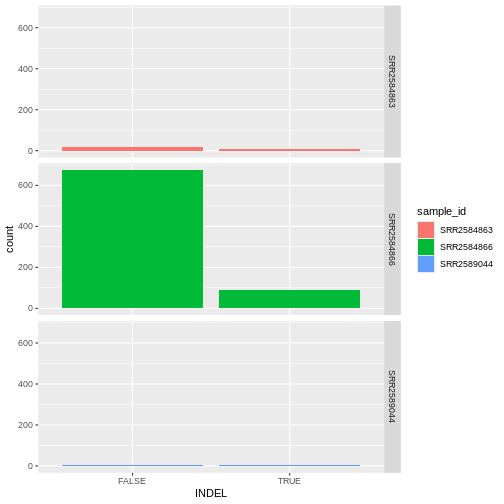
We can create barplots using the geom_bar
geom. Let’s make a barplot showing the number of variants for each
sample that are indels.
R
ggplot(data = variants, aes(x = INDEL, fill = sample_id)) +
geom_bar() +
facet_grid(sample_id ~ .)
Challenge
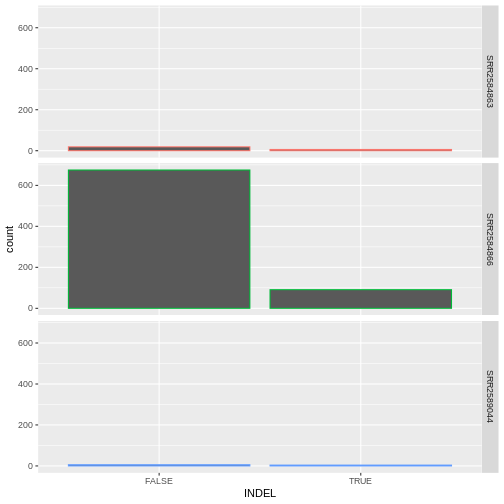
- From the previus plot, we realize we don’t need the legend since we
already have the sample_id labels on the individual plot facets. Use the
help file for
geom_barand any other online resources you want to use to remove the legend from the plot.
R
ggplot(data = variants, aes(x = INDEL, color = sample_id)) +
geom_bar(show.legend = F) +
facet_grid(sample_id ~ .)
Density
We can create density plots using the geom_density
geom that shows the distribution of of a variable in the dataset. Let’s
plot the distribution of QUAL
R
ggplot(data = variants, aes(x = DP)) +
geom_density()
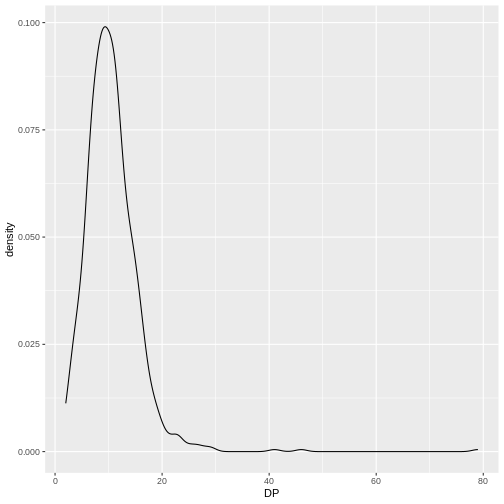
This plot tells us that the most of frequent DP (read
depth) for the variants is about 10 reads.
Challenge
Use geom_density
to plot the distribution of QUAL with a different fill for
each sample. Use a white background for the plot.
R
ggplot(data = variants, aes(x = QUAL, fill = sample_id)) +
geom_density(alpha = 0.3) +
theme_bw()
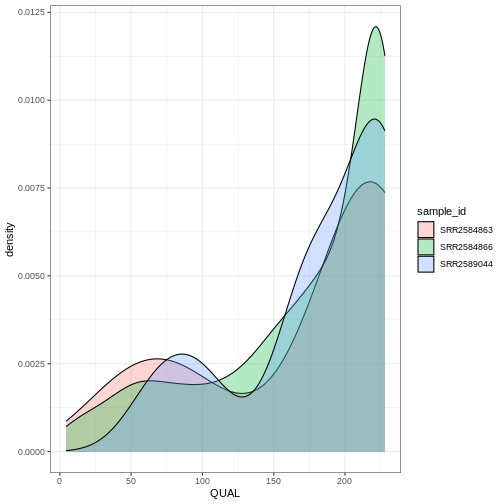
ggplot2 themes
In addition to theme_bw(),
which changes the plot background to white,
ggplot2 comes with several other themes
which can be useful to quickly change the look of your visualization.
The complete list of themes is available at https://ggplot2.tidyverse.org/reference/ggtheme.html.
theme_minimal() and theme_light() are popular,
and theme_void() can be useful as a starting point to
create a new hand-crafted theme.
The ggthemes
package provides a wide variety of options (including Microsoft Excel,
old
and new).
The ggplot2
extensions website provides a list of packages that extend the
capabilities of ggplot2, including
additional themes.
Challenge
With all of this information in hand, please take another five
minutes to either improve one of the plots generated in this exercise or
create a beautiful graph of your own. Use the RStudio ggplot2
cheat sheet for inspiration. Here are some ideas:
- See if you can change the size or shape of the plotting symbol.
- Can you find a way to change the name of the legend? What about its labels?
- Try using a different color palette (see the Cookbook for R).
More ggplot2 Plots
ggplot2 offers many more informative
and beautiful plots (geoms) of interest for biologists
(although not covered in this lesson) that are worth exploring, such
as
-
geom_tile(), for heatmaps, -
geom_jitter(), for strip charts, and -
geom_violin(), for violin plots
Resources
- ggplot2: Elegant Graphics for Data Analysis (online version)
- The Grammar of Graphics (Statistics and Computing)
- Data Visualization: A Practical Introduction (online version)
- The R Graph Gallery (the book)
- ggplot2 is a powerful tool for high-quality plots
- ggplot2 provides a flexible and readable grammar to build plots
